UGENE is free open-source cross-platform set of integrated bioinformatics software.
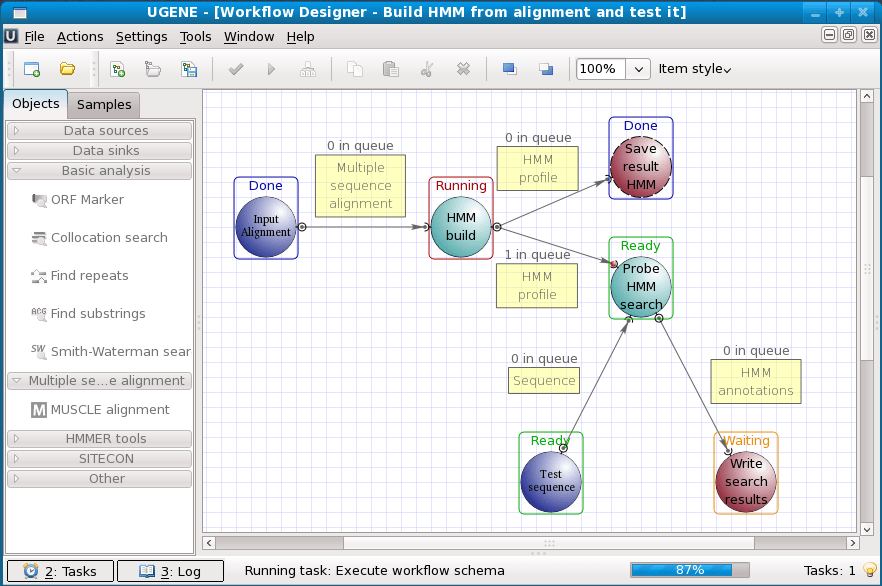
It integrates dozens of well-known biological tools and algorithms, providing both graphical user and command line interfaces. Using UGENE Workflow Designer one can arrange the required tools and algorithms into a workflow schema.
In order to provide maximum possible performance UGENE utilizes multicore CPUs and GPUs to optimize some of its computational routines. Another way to speed up computations is to use Amazon EC2 cloud resources.
Key Features
- Graphical User interface offering:
- Visual and interactive genome browsing including circular plasmid view.
- Multiple alignment editor.
- Chromatograms visualization.
- 3D viewer for files in PDB and MMDB formats with anaglyph stereo mode support.
- Phylogenetic tree viewer.
- Dot plot visualization.
- Query designer enables search for complex annotation patterns.
- Easy to use workflow designer for custom computational workflow.
- Multiple sequence alignment using MUSCLE 3, KAlign, Clustal.
- HMM profiles build and search, based on the source of HMMER 2 and HMMER 3.
- PCR Primers design using Primer 3.
- Cloning in silico capabilities.
- Queries to NCBI BLAST and CDD databases.
- Protein secondary structure prediction using GOR IV and PSIPRED.
- Phylogenetic analysis with Phylip.
- Search for restriction enzymes and integration with REBASE.
- Extremely fast repeat finder.
- DNA reference assembly using Bowtie.
- Search for transcription factor binding sites using SITECON.
- Protein back translation.
- ORF finder.
- Complete Smith-Waterman algorithm implementation.
- Support for external tools: BLAST+, MAFFT, T-Coffee.
- Assembly Browser — a shiny BAM viewer.
- Biostruct3D viewer.
- Complete support of modern multicore processors and SSE instructions.
- Out of the box support of modern GPUs using NVIDIA CUDA and ATI Stream.
- Support for cloud computing: running computational tasks on Amazon EC2 cloud.
- Support for supercomputers and distributed computing.
Website: ugene.net
Support: Manual
Developer: Unipro
License: GNU General Public License v2.0
UGENE is written in C++. Learn C++ with our recommended free books and free tutorials.
Related Software
| Bioinformatics Tools | |
|---|---|
| Bioconductor | Analysis and comprehension of high-throughput genomic data |
| Biopython | Tools for biological computation written in Python |
| UGENE | Set of integrated bioinformatics software |
| BioPerl | Perl tools for computational molecular biology |
| GROMACS | Versatile package to perform molecular dynamics |
| IGV | High-performance visualization genome browser tool |
| GATK | Genomic analysis toolkit focused on variant discovery |
| BioJava | Provides Java tools for processing biological data |
| InterMine | Integrate biological data sources |
| bedtools | Powerful toolset for genome arithmetic |
| EMBOSS | The European Molecular Biology Open Software Suite |
| BLAST | Algorithm for comparing primary biological sequence information |
| Galaxy | Web-based platform for data-intensive computational research |
| minimap2 | Versatile sequence alignment program |
| Jalview | Multiple sequence alignment editing, visualisation and analysis |
| samtools | Manipulate next-generation sequencing data |
| BCFtools | Variant calling and manipulating files in the Variant Call Format |
| FastQC | Quality control tool for high throughput sequence data |
| SPAdes | Versatile toolkit for assembling and analysing sequencing data |
| GenomeTools | Collection of bioinformatics tools |
| AliView | Alignment viewer and editor |
| mothur | Analyze microbial communities |
| Bandage | Visualising de novo assembly graphs |
| cramino | BAM/CRAM quality evaluation |
| abPOA | Adaptive banded Partial Order Alignment |
| Taverna Workbench | For designing and executing bioinformatics workflows |
| geWorkbench | Software platform for integrated genomic data analysis |
| Bioclipse | Rich-client platform chemistry and biology workbench |
Read our verdict in the software roundup.
| Desktop Genome Browsers | |
|---|---|
| IGV | High-performance visualization genome browser tool |
| IGB | Integrated Genome Browser |
| JBrowse | Fast, scalable genome browser |
| Tablet | Graphical viewer for sequence assemblies and alignments |
| UGENE | Set of integrated bioinformatics software |
| Artemis | Genome viewer and annotation tool |
| Apollo | Instantaneous, collaborative genomic annotation editor |
| CutePeaks | Cross platform Sanger Trace file viewer |
| Genome Workbench | Integrated tools for studying and analyzing genetic data |
Read our verdict in the software roundup.
 Explore our comprehensive directory of recommended free and open source software. Our carefully curated collection spans every major software category. Explore our comprehensive directory of recommended free and open source software. Our carefully curated collection spans every major software category.This directory is part of our ongoing series of informative articles for Linux enthusiasts. It features hundreds of detailed reviews, along with open source alternatives to proprietary solutions from major corporations such as Google, Microsoft, Apple, Adobe, IBM, Cisco, Oracle, and Autodesk. You’ll also find interesting projects to try, hardware coverage, free programming books and tutorials, and much more. Discovered a useful open source Linux program that we haven’t covered yet? Let us know by completing this form. |

