geWorkbench (genomics Workbench) is a Java-based open-source platform for integrated genomics. Using a component architecture it allows individually developed plug-ins to be configured into complex bioinformatic applications.
At present there are more than 70 available plug-ins supporting the visualization and analysis of gene expression and sequence data.
geWorkbench is the Bioinformatics platform of MAGNet, the National Center for the Multi-scale Analysis of Genomic and Cellular Networks, one of the 8 National Centers for Biomedical Computing.
Key Features
- Computational analysis tools such as t-test, hierarchical clustering, self-organizing maps, regulatory network reconstruction, BLAST searches, pattern-motif discovery, protein structure prediction, structure-based protein annotation, etc.
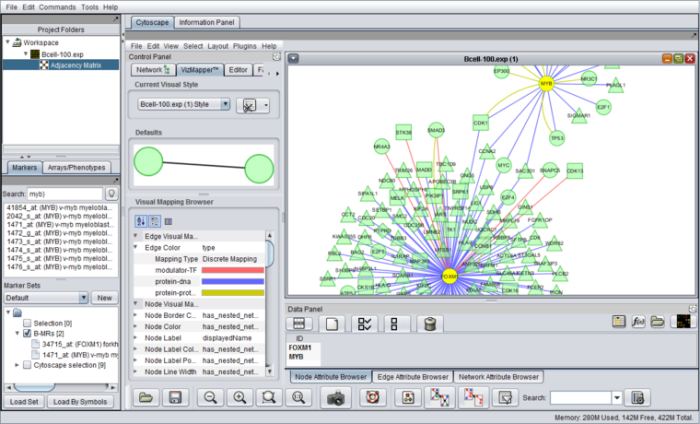
- Visualization of gene expression (heatmaps, volcano plot), molecular interaction networks (through Cytoscape), protein sequence and protein structure data (e.g., MarkUs).
- Integration of gene and pathway annotation information from curated sources as well as through Gene Ontology enrichment analysis.
- Component integration through platform management of inputs and outputs. Among data that can be shared between components are expression datasets, interaction networks, sample and marker (gene) sets and sequences.
- Dataset history tracking – complete record of data sets used and input settings.
- Integration with 3rd party tools such as Genepattern, Cytoscape, and Genomespace.
- Provides an environment which supports moving from one data type to another in a seamless fashion, e.g. from gene expression to sequences to patterns.
- Provides access to a variety of external data sources, including:
- Microarray gene expression repositories (caArray).
- BLAST (NCBI).
- Gene annotation pages (via bioDBNet).
- Protein and DNA sequence retrieval (UC Santa Cruz and EBI).
- Pathway diagrams (BioCarta).
- Provides a gateway to several computational services currently hosted on Columbia servers and clusters, including:
-
- Pattern Discovery.
- Pudge – protein structure modeling.
- SkyBase – database of molecular models.
-
Specific types of data supported include:
- Microarray Gene Expression:
- GEO Soft: Series, Series Matrix, and Annotated Matrix (GDS).
- MAGE-TAB data matrix.
- Affymetrix GCOS/MAS5.
- Matrix format (geWorkbench).
- Tab-delimited (e.g. RMAExpress).
- GenePix.
- Microarray Gene Expression Annotation file support:
- Affymetrix 3′ Expression.
- Affymetrix WT Gene/Exon ST (transcript-level) including Gene Array 1.0/2.0 ST and Exon 1.0 ST.
- DNA and Protein Sequences:
- FASTA.
- Pathways:
- BioCarta.
- Molecular structure – prediction, annotation and display.
- Sequence Patterns:
- Regular Expressions.
- Gene Ontology.
- Regulatory Networks.
Website: github.com/floratos-lab/geworkbench-core
Support:
Developer: Columbia University, First Genetic Trust National Cancer Institute
License: BSD-like
geWorkbench is written in Java. Learn Java with our recommended free books and free tutorials.
Related Software
| Bioinformatics Tools | |
|---|---|
| Bioconductor | Analysis and comprehension of high-throughput genomic data |
| Biopython | Tools for biological computation written in Python |
| UGENE | Set of integrated bioinformatics software |
| BioPerl | Perl tools for computational molecular biology |
| GROMACS | Versatile package to perform molecular dynamics |
| IGV | High-performance visualization genome browser tool |
| GATK | Genomic analysis toolkit focused on variant discovery |
| BioJava | Provides Java tools for processing biological data |
| InterMine | Integrate biological data sources |
| bedtools | Powerful toolset for genome arithmetic |
| EMBOSS | The European Molecular Biology Open Software Suite |
| BLAST | Algorithm for comparing primary biological sequence information |
| Galaxy | Web-based platform for data-intensive computational research |
| minimap2 | Versatile sequence alignment program |
| Jalview | Multiple sequence alignment editing, visualisation and analysis |
| samtools | Manipulate next-generation sequencing data |
| BCFtools | Variant calling and manipulating files in the Variant Call Format |
| FastQC | Quality control tool for high throughput sequence data |
| SPAdes | Versatile toolkit for assembling and analysing sequencing data |
| GenomeTools | Collection of bioinformatics tools |
| AliView | Alignment viewer and editor |
| mothur | Analyze microbial communities |
| Bandage | Visualising de novo assembly graphs |
| cramino | BAM/CRAM quality evaluation |
| abPOA | Adaptive banded Partial Order Alignment |
| Taverna Workbench | For designing and executing bioinformatics workflows |
| geWorkbench | Software platform for integrated genomic data analysis |
| Bioclipse | Rich-client platform chemistry and biology workbench |
Read our verdict in the software roundup.
 Explore our comprehensive directory of recommended free and open source software. Our carefully curated collection spans every major software category. Explore our comprehensive directory of recommended free and open source software. Our carefully curated collection spans every major software category.This directory is part of our ongoing series of informative articles for Linux enthusiasts. It features hundreds of detailed reviews, along with open source alternatives to proprietary solutions from major corporations such as Google, Microsoft, Apple, Adobe, IBM, Cisco, Oracle, and Autodesk. You’ll also find interesting projects to try, hardware coverage, free programming books and tutorials, and much more. Discovered a useful open source Linux program that we haven’t covered yet? Let us know by completing this form. |

