The GenomeTools genome analysis system is a collection of bioinformatics tools (in the realm of genome informatics) combined into a single binary named gt.
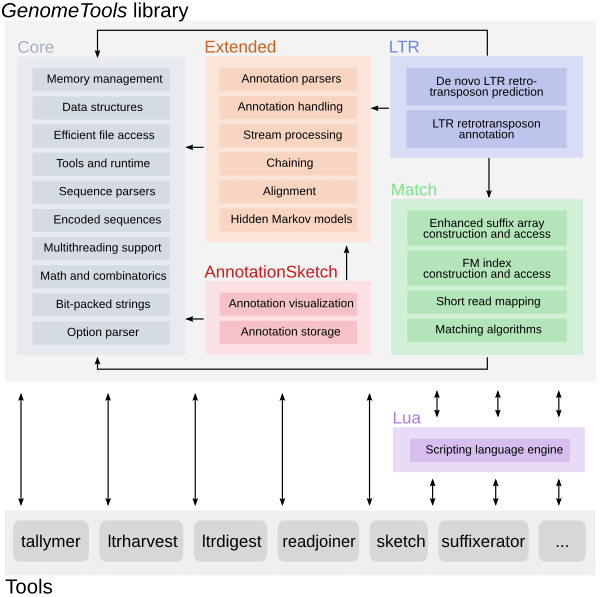
It is based on a C library named libgenometools which contains a wide variety of classes for efficient and convenient implementation of sequence and annotation processing software.
GenomeTools has been designed to run on every POSIX compliant UNIX system — for example Linux or macOS. A reduced Windows version is also available.
This is free and open source software.
Some of the tools are:
- LTRharvest, an efficient and flexible software tool for de novo detection of LTR retrotransposons.
- Tallymer, a collection of flexible and memory-efficient programs for k-mer counting and indexing of large sequence sets.
- uniquesub, a program for computing minimum unique substrings.
- AnnotationSketch, a library for drawing genome annotations.
- LTRdigest, a software tool for automated annotation of internal features of LTR retrotransposons.
- MetaGenomeThreader, a software to predict genes, such as PCS’s (predicted coding sequences) in sequences of metagenome projects.
- GtEncseq, a compressed biosequence representation with many features.
- Readjoiner, a sequence assembler based on the assembly string graph framework.
Website: genometools.org
Support: GitHub Code Repository
Developer: Broad Institute, Inc.
License: Apache License 2.0
GenomeTools is written in C. Learn C with our recommended free books and free tutorials.
Related Software
| Bioinformatics Tools | |
|---|---|
| Bioconductor | Analysis and comprehension of high-throughput genomic data |
| Biopython | Tools for biological computation written in Python |
| UGENE | Set of integrated bioinformatics software |
| BioPerl | Perl tools for computational molecular biology |
| GROMACS | Versatile package to perform molecular dynamics |
| IGV | High-performance visualization genome browser tool |
| GATK | Genomic analysis toolkit focused on variant discovery |
| BioJava | Provides Java tools for processing biological data |
| InterMine | Integrate biological data sources |
| bedtools | Powerful toolset for genome arithmetic |
| EMBOSS | The European Molecular Biology Open Software Suite |
| BLAST | Algorithm for comparing primary biological sequence information |
| Galaxy | Web-based platform for data-intensive computational research |
| minimap2 | Versatile sequence alignment program |
| Jalview | Multiple sequence alignment editing, visualisation and analysis |
| samtools | Manipulate next-generation sequencing data |
| BCFtools | Variant calling and manipulating files in the Variant Call Format |
| FastQC | Quality control tool for high throughput sequence data |
| SPAdes | Versatile toolkit for assembling and analysing sequencing data |
| GenomeTools | Collection of bioinformatics tools |
| AliView | Alignment viewer and editor |
| mothur | Analyze microbial communities |
| Bandage | Visualising de novo assembly graphs |
| cramino | BAM/CRAM quality evaluation |
| abPOA | Adaptive banded Partial Order Alignment |
| Taverna Workbench | For designing and executing bioinformatics workflows |
| geWorkbench | Software platform for integrated genomic data analysis |
| Bioclipse | Rich-client platform chemistry and biology workbench |
Read our verdict in the software roundup.
 Explore our comprehensive directory of recommended free and open source software. Our carefully curated collection spans every major software category. Explore our comprehensive directory of recommended free and open source software. Our carefully curated collection spans every major software category.This directory is part of our ongoing series of informative articles for Linux enthusiasts. It features hundreds of detailed reviews, along with open source alternatives to proprietary solutions from major corporations such as Google, Microsoft, Apple, Adobe, IBM, Cisco, Oracle, and Autodesk. You’ll also find interesting projects to try, hardware coverage, free programming books and tutorials, and much more. Discovered a useful open source Linux program that we haven’t covered yet? Let us know by completing this form. |

