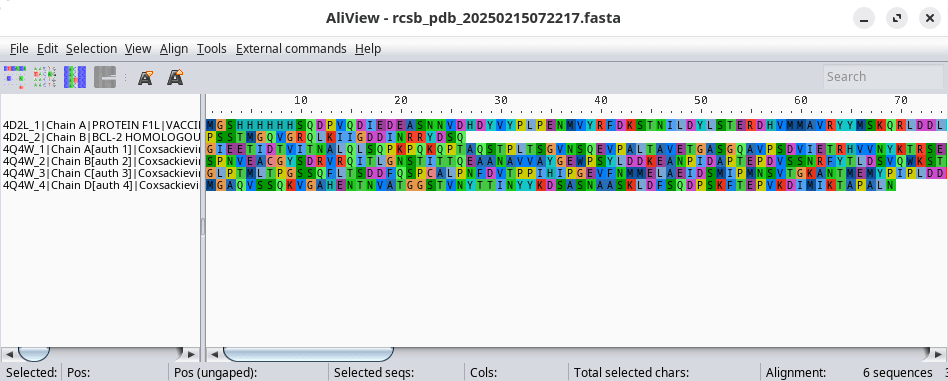
AliView is an alignment viewer and editor of DNA / aminoacid sequences. It aims to be one of the fastest and most intuitive to use.
The general idea when designing this program has always been usability and speed, all new functions are optimized so they do not affect the general performance and capability to work swiftly with large alignments.
A need to easily sort, view, remove, edit and merge sequences from large transcriptome datasets initiated the work with the program.
Key Features
- Supports Fasta, Nexus, Phylip, Clustal or MSF-format (unlimited file sizes).
edit (manually). - Align, add and align automatically (with MUSCLE or MAFFT or any other aligner of your choice).
- Find degenerate primers in conserved regions in an alignment of mixed species.
- On the fly translation of nucleotides to amino acids.
- Various visual cues to highlight consensus characters or characters deviating from the consensus.
- Simple copy/paste/drop/remove of sequences/files.
- Very simple to use “external interface” that lets you invoke your other favorite programs (you could for example automatically have the alignment sent to FastTree and then automatically opened in FigTree).
- Cross-platform support – runs under Linux, macOS, and Windows.
Website: www.ormbunkar.se/aliview
Support: GitHub Code Repository
Developer: Department of Systematic Biology (Uppsala University)
License: GNU General Public License v3.0
AliView is written in Java. Learn Java with our recommended free books and free tutorials.
Related Software
| Bioinformatics Tools | |
|---|---|
| Bioconductor | Analysis and comprehension of high-throughput genomic data |
| Biopython | Tools for biological computation written in Python |
| UGENE | Set of integrated bioinformatics software |
| BioPerl | Perl tools for computational molecular biology |
| GROMACS | Versatile package to perform molecular dynamics |
| IGV | High-performance visualization genome browser tool |
| GATK | Genomic analysis toolkit focused on variant discovery |
| BioJava | Provides Java tools for processing biological data |
| InterMine | Integrate biological data sources |
| bedtools | Powerful toolset for genome arithmetic |
| EMBOSS | The European Molecular Biology Open Software Suite |
| BLAST | Algorithm for comparing primary biological sequence information |
| Galaxy | Web-based platform for data-intensive computational research |
| minimap2 | Versatile sequence alignment program |
| Jalview | Multiple sequence alignment editing, visualisation and analysis |
| samtools | Manipulate next-generation sequencing data |
| BCFtools | Variant calling and manipulating files in the Variant Call Format |
| FastQC | Quality control tool for high throughput sequence data |
| SPAdes | Versatile toolkit for assembling and analysing sequencing data |
| GenomeTools | Collection of bioinformatics tools |
| AliView | Alignment viewer and editor |
| mothur | Analyze microbial communities |
| Bandage | Visualising de novo assembly graphs |
| cramino | BAM/CRAM quality evaluation |
| abPOA | Adaptive banded Partial Order Alignment |
| Taverna Workbench | For designing and executing bioinformatics workflows |
| geWorkbench | Software platform for integrated genomic data analysis |
| Bioclipse | Rich-client platform chemistry and biology workbench |
Read our verdict in the software roundup.
 Explore our comprehensive directory of recommended free and open source software. Our carefully curated collection spans every major software category. Explore our comprehensive directory of recommended free and open source software. Our carefully curated collection spans every major software category.This directory is part of our ongoing series of informative articles for Linux enthusiasts. It features hundreds of detailed reviews, along with open source alternatives to proprietary solutions from major corporations such as Google, Microsoft, Apple, Adobe, IBM, Cisco, Oracle, and Autodesk. You’ll also find interesting projects to try, hardware coverage, free programming books and tutorials, and much more. Discovered a useful open source Linux program that we haven’t covered yet? Let us know by completing this form. |

