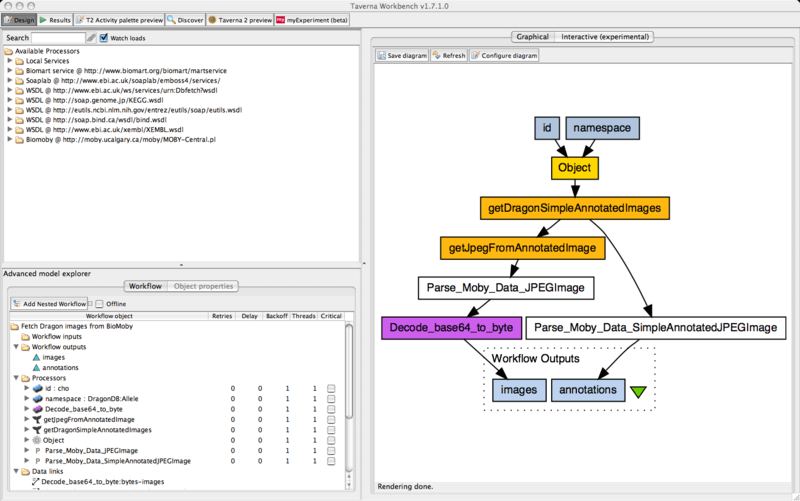
The Taverna workbench is an open source software tool for designing and executing bioinformatics workflows created by the myGrid project.
Taverna allows users to integrate many different software components, including SOAP or REST Web services, such as those provided by the National Center for Biotechnology Information, the European Bioinformatics Institute, the DNA Databank of Japan (DDBJ), SoapLab, BioMOBY and EMBOSS. The set of available services is not finite and users can import new service descriptions into the Taverna Workbench.
The Taverna suite is written in Java and includes the Taverna Engine (used for enacting workflows) that powers both the Taverna Workbench (the desktop client application) and the Taverna Server (which allows remote execution of workflows). Taverna is also available as a Command Line Tool for a quick execution of workflows from a terminal.
Taverna is used by users in many domains, such as bioinformatics, cheminformatics, medicine, astronomy, social science and music.
Key Features
- Fully featured, extensible and scalable scientific Workflow Management System:
- Tabs for finding, designing and executing workflows.
- Fully graphical workflow design.
- Drag and drop workflow components.
- Comprehensive undo/redo.
- Built-in help facility.
- Annotations for describing workflows, services, inputs, outputs.
- Workflow validation and debugging.
- Execute and debug workflows:
- Execute workflows.
- Remember previously used workflow inputs.
- Save workflow input values used to a file.
- Load workflow input values from a file.
- Pipelining and streaming of data.
- Implicit iteration of service calls.
- Conditional and repeated calling of services.
- Customizable looping over a service.
- Failover and retry of service calling.
- Parallel execution and configurable number of concurrent threads.
- Improved error handling and reporting for debugging during run time.
- Monitor workflow execution.
- Pause/resume or cancel workflow execution.
- Manage previous runs and workflow results.
- View intermediate results and debug workflows at run time.
- Filter and save intermediate and final workflow results.
- Available as a desktop Workbench, from a command line or remotely as a Server.
- Access to local and remote resources and analysis tools, Web and grid services; 3500+ service s available on start up.
- Not restricted to predetermined services – rapid incorporation of new services without coding.
- Extensible service plug-in architecture for adding new service type.
- Excel and csv spreadsheet support.
- Invoke general SOAP/WSDL or REST Web services, and more specific SADI, BioMart, BioMoby and SoapLab Web services.
- Invoke R statistical services.
- Invoke local Java code.
- Invoke external tools on remote machines (via ssh), D.
- Do XPath and other text manipulation.
- Import a spreadsheet and include sub-workflows.
- Search for services described in BioCatalogue to include within workflows.
- Search for workflows on myExperiment. You can download, modify and run the workflows discovered on myExperiment from within the Taverna Workbench.
- Plugins to extend functionality.
Website: taverna.incubator.apache.org
Support: Documentation
Developer: myGrid
License: GNU LGPL 2.1
Taverna is written in Java. Learn Java with our recommended free books and free tutorials.
Related Software
| Bioinformatics Tools | |
|---|---|
| Bioconductor | Analysis and comprehension of high-throughput genomic data |
| Biopython | Tools for biological computation written in Python |
| UGENE | Set of integrated bioinformatics software |
| BioPerl | Perl tools for computational molecular biology |
| GROMACS | Versatile package to perform molecular dynamics |
| IGV | High-performance visualization genome browser tool |
| GATK | Genomic analysis toolkit focused on variant discovery |
| BioJava | Provides Java tools for processing biological data |
| InterMine | Integrate biological data sources |
| bedtools | Powerful toolset for genome arithmetic |
| EMBOSS | The European Molecular Biology Open Software Suite |
| BLAST | Algorithm for comparing primary biological sequence information |
| Galaxy | Web-based platform for data-intensive computational research |
| minimap2 | Versatile sequence alignment program |
| Jalview | Multiple sequence alignment editing, visualisation and analysis |
| samtools | Manipulate next-generation sequencing data |
| BCFtools | Variant calling and manipulating files in the Variant Call Format |
| FastQC | Quality control tool for high throughput sequence data |
| SPAdes | Versatile toolkit for assembling and analysing sequencing data |
| GenomeTools | Collection of bioinformatics tools |
| AliView | Alignment viewer and editor |
| mothur | Analyze microbial communities |
| Bandage | Visualising de novo assembly graphs |
| cramino | BAM/CRAM quality evaluation |
| abPOA | Adaptive banded Partial Order Alignment |
| Taverna Workbench | For designing and executing bioinformatics workflows |
| geWorkbench | Software platform for integrated genomic data analysis |
| Bioclipse | Rich-client platform chemistry and biology workbench |
Read our verdict in the software roundup.
 Explore our comprehensive directory of recommended free and open source software. Our carefully curated collection spans every major software category. Explore our comprehensive directory of recommended free and open source software. Our carefully curated collection spans every major software category.This directory is part of our ongoing series of informative articles for Linux enthusiasts. It features hundreds of detailed reviews, along with open source alternatives to proprietary solutions from major corporations such as Google, Microsoft, Apple, Adobe, IBM, Cisco, Oracle, and Autodesk. You’ll also find interesting projects to try, hardware coverage, free programming books and tutorials, and much more. Discovered a useful open source Linux program that we haven’t covered yet? Let us know by completing this form. |

