The Integrative Genomics Viewer (IGV) is a high-performance visualization tool for interactive exploration of large, integrated genomic datasets. It supports a wide variety of data types, including array-based and next-generation sequence data, and genomic annotations.
Navigation through a dataset is similar to Google Maps, allowing the user to zoom and pan seamlessly across the genome at any level of detail from whole-genome to base pair.
IGV’s scalable architecture makes it well suited for genome-wide exploration of next-generation sequencing (NGS) datasets, including both basic aligned read data as well as derived results, such as read coverage.
Key Features
- Flexible integration of a wide range of genomic data types including aligned sequence reads, mutations, copy number, RNAi screens, gene expression, methylation, and genomic annotation.
- Use of efficient, multi-resolution file formats to enable real-time exploration of arbitrarily large datasets over all resolution scales, while consuming minimal resources on the client computer.
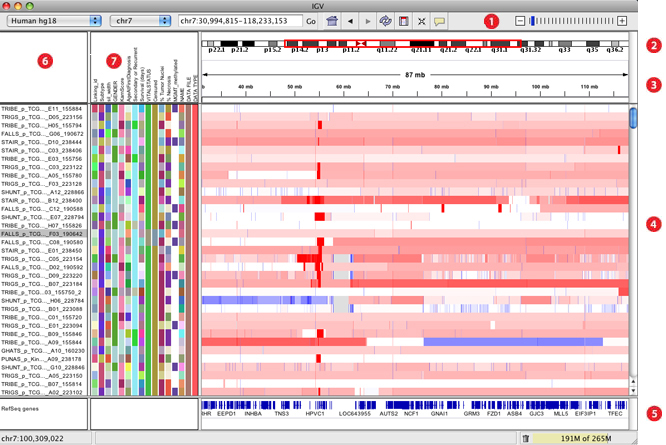
- Supports concurrent visualization of diverse data types across hundreds, and up to thousands of samples, and correlation of these integrated datasets with clinical and phenotypic variables
- Datasets can be loaded from local or remote sources, including cloud-based resources, enabling investigators to view their own genomic datasets alongside publicly available data from, for example, The Cancer Genome Atlas (TCGA, 1000 Genomes (www.1000genomes.org/), and ENCODE (www.genome.gov/10005107) projects. In addition, IGV allows collaborators to load and share data locally or remotely over the Web.
- Cross-platform support – runs under Linux, Mac OS X, Windows, and any other operating system that supports Java 8 or 11.
Website: igv.org/doc/desktop
Support: User Guide, GitHub Code Repository
Developer: Broad Institute and Regents of the University of California
License: MIT License
IGV is written in Java. Learn Java with our recommended free books and free tutorials.
Related Software
| Bioinformatics Tools | |
|---|---|
| Bioconductor | Analysis and comprehension of high-throughput genomic data |
| Biopython | Tools for biological computation written in Python |
| UGENE | Set of integrated bioinformatics software |
| BioPerl | Perl tools for computational molecular biology |
| GROMACS | Versatile package to perform molecular dynamics |
| IGV | High-performance visualization genome browser tool |
| GATK | Genomic analysis toolkit focused on variant discovery |
| BioJava | Provides Java tools for processing biological data |
| InterMine | Integrate biological data sources |
| bedtools | Powerful toolset for genome arithmetic |
| EMBOSS | The European Molecular Biology Open Software Suite |
| BLAST | Algorithm for comparing primary biological sequence information |
| Galaxy | Web-based platform for data-intensive computational research |
| minimap2 | Versatile sequence alignment program |
| Jalview | Multiple sequence alignment editing, visualisation and analysis |
| samtools | Manipulate next-generation sequencing data |
| BCFtools | Variant calling and manipulating files in the Variant Call Format |
| FastQC | Quality control tool for high throughput sequence data |
| SPAdes | Versatile toolkit for assembling and analysing sequencing data |
| GenomeTools | Collection of bioinformatics tools |
| AliView | Alignment viewer and editor |
| mothur | Analyze microbial communities |
| Bandage | Visualising de novo assembly graphs |
| cramino | BAM/CRAM quality evaluation |
| abPOA | Adaptive banded Partial Order Alignment |
| Taverna Workbench | For designing and executing bioinformatics workflows |
| geWorkbench | Software platform for integrated genomic data analysis |
| Bioclipse | Rich-client platform chemistry and biology workbench |
Read our verdict in the software roundup.
| Desktop Genome Browsers | |
|---|---|
| IGV | High-performance visualization genome browser tool |
| IGB | Integrated Genome Browser |
| JBrowse | Fast, scalable genome browser |
| Tablet | Graphical viewer for sequence assemblies and alignments |
| UGENE | Set of integrated bioinformatics software |
| Artemis | Genome viewer and annotation tool |
| Apollo | Instantaneous, collaborative genomic annotation editor |
| CutePeaks | Cross platform Sanger Trace file viewer |
| Genome Workbench | Integrated tools for studying and analyzing genetic data |
Read our verdict in the software roundup.
 Explore our comprehensive directory of recommended free and open source software. Our carefully curated collection spans every major software category. Explore our comprehensive directory of recommended free and open source software. Our carefully curated collection spans every major software category.This directory is part of our ongoing series of informative articles for Linux enthusiasts. It features hundreds of detailed reviews, along with open source alternatives to proprietary solutions from major corporations such as Google, Microsoft, Apple, Adobe, IBM, Cisco, Oracle, and Autodesk. You’ll also find interesting projects to try, hardware coverage, free programming books and tutorials, and much more. Discovered a useful open source Linux program that we haven’t covered yet? Let us know by completing this form. |

