Jalview is a cross-platform program for multiple sequence alignment editing, visualisation and analysis. Use it to view and edit sequence alignments, analyse them with phylogenetic trees and principal components analysis (PCA) plots and explore molecular structures and annotation.
The Jalview Desktop is an application you can launch from the web or install locally. It is designed for the kind of in-depth sequence analysis necessary when exploring new protein or RNA sequence families to understand how their sequence relates to structure and function.
The Desktop can access sequence, alignment, 3D structure and annotation databases, and web services for multiple alignment, protein disorder and secondary structure prediction and functional analysis.
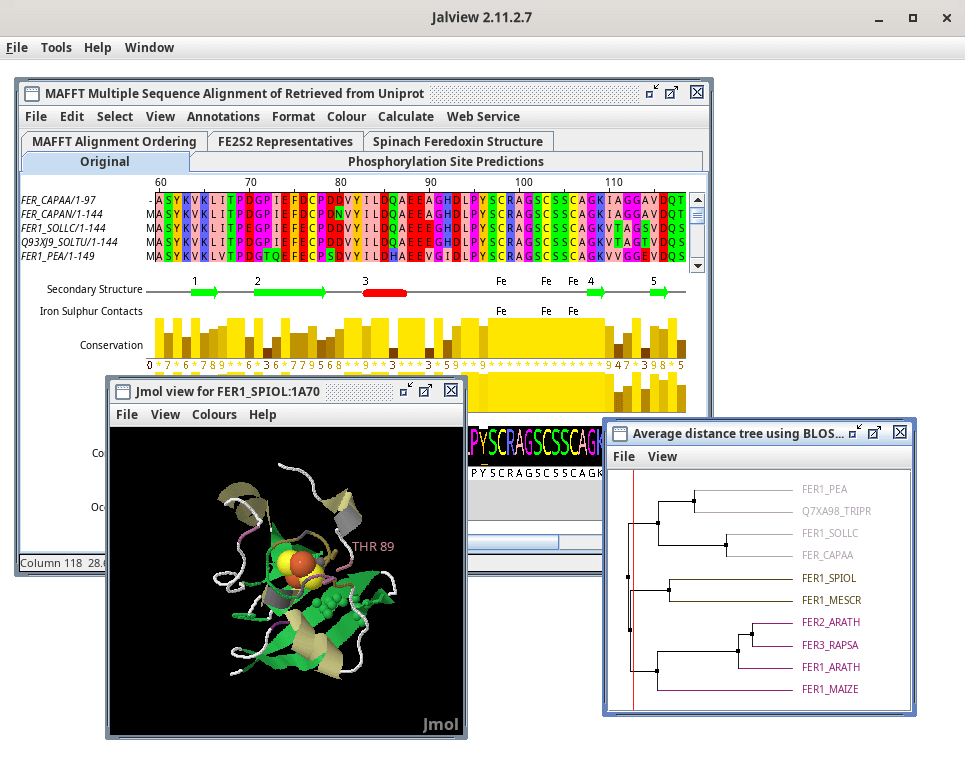
Jalview is able to link views of 3D structures to multiple sequence alignments. Jmol is integrated with Jalview for this, but you can also configure Jalview to use Chimera, ChimeraX and Pymol for 3D structure visualisation.
Alignments can also be linked to RNA secondary structure visualisations with VARNA, also included in the Jalview Desktop.
Website: www.jalview.org
Support:
Developer: Managed by The Barton Group, University of Dundee, UK
License: GNU General Public License v3.0
Jalview is written in Java. Learn Java with our recommended free books and free tutorials.
Related Software
| Bioinformatics Tools | |
|---|---|
| Bioconductor | Analysis and comprehension of high-throughput genomic data |
| Biopython | Tools for biological computation written in Python |
| UGENE | Set of integrated bioinformatics software |
| BioPerl | Perl tools for computational molecular biology |
| GROMACS | Versatile package to perform molecular dynamics |
| IGV | High-performance visualization genome browser tool |
| GATK | Genomic analysis toolkit focused on variant discovery |
| BioJava | Provides Java tools for processing biological data |
| InterMine | Integrate biological data sources |
| bedtools | Powerful toolset for genome arithmetic |
| EMBOSS | The European Molecular Biology Open Software Suite |
| BLAST | Algorithm for comparing primary biological sequence information |
| Galaxy | Web-based platform for data-intensive computational research |
| minimap2 | Versatile sequence alignment program |
| Jalview | Multiple sequence alignment editing, visualisation and analysis |
| samtools | Manipulate next-generation sequencing data |
| BCFtools | Variant calling and manipulating files in the Variant Call Format |
| FastQC | Quality control tool for high throughput sequence data |
| SPAdes | Versatile toolkit for assembling and analysing sequencing data |
| GenomeTools | Collection of bioinformatics tools |
| AliView | Alignment viewer and editor |
| mothur | Analyze microbial communities |
| Bandage | Visualising de novo assembly graphs |
| cramino | BAM/CRAM quality evaluation |
| abPOA | Adaptive banded Partial Order Alignment |
| Taverna Workbench | For designing and executing bioinformatics workflows |
| geWorkbench | Software platform for integrated genomic data analysis |
| Bioclipse | Rich-client platform chemistry and biology workbench |
Read our verdict in the software roundup.
 Explore our comprehensive directory of recommended free and open source software. Our carefully curated collection spans every major software category. Explore our comprehensive directory of recommended free and open source software. Our carefully curated collection spans every major software category.This directory is part of our ongoing series of informative articles for Linux enthusiasts. It features hundreds of detailed reviews, along with open source alternatives to proprietary solutions from major corporations such as Google, Microsoft, Apple, Adobe, IBM, Cisco, Oracle, and Autodesk. You’ll also find interesting projects to try, hardware coverage, free programming books and tutorials, and much more. Discovered a useful open source Linux program that we haven’t covered yet? Let us know by completing this form. |

