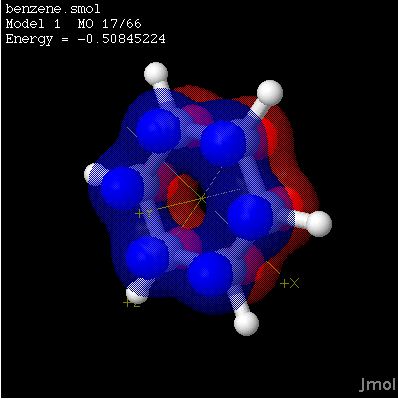
Jmol is an open-source Java viewer for three-dimensional chemical structures with features for chemicals, crystals, materials and biomolecules.
Jmol returns a 3D representation of a molecule that may be used as a teaching tool, or for research e.g. in chemistry and biochemistry.
The software is for students, educators, and researchers in chemistry, biochemistry, physics, and materials science.
Key Features
- Applet, Application, and Systems Integration Component:
- The JmolApplet is a web browser applet that can be integrated into web pages. It is ideal for development of web-based courseware and web-accessible chemical databases. The JmolApplet provides an upgrade path for users of the Chime plug-in.
- The Jmol application is a standalone Java application that runs on the desktop.
- The JmolViewer can be integrated as a component into other Java applications.
- High-performance 3D rendering with no hardware requirements.
- Supports a wide range of molecular file formats:
- MOD – MDL / Elsevier / Symyx structure (classic version V2000).
- V3000 – MDL / Elsevier / Symyx structure (new version V3000).
- SDF – MDL / Elsevier / Symyx structure (multiple models).
- CTFile – MDL / Elsevier / Symyx chemical table (generic).
- CIF – Crystallographic Information File – standard from the International Union of Crystallography.
- mmCIF – Macromolecular Crystallographic Information File – standard from the International Union of Crystallography.
- CML – Chemical Markup Language.
- PDB – Protein Data Bank – Research Collaboratory for Structural Bioinformatics.
- XYZ – XYZ format, XMol file – Minnesota Supercomputer Institute.
- XYZ+vib – XYZ format with added vibrational vector information.
- XYZ-FAH – XYZ format for Folding@home.
- MOL2 – Sybyl, Tripos.
- Alchemy – Tripos.
- CSF – Fujitsu CAChe chemical structure, now Fujitsu Sygress.
- GAMESS – General Atomic and Molecular Electronic Structure System output (both US and UK variants) – Gordon Research Group, Iowa State University.
- Gaussian – Gaussian 94/98/03 output – Gaussian, Inc.
- Cube – Gaussian, Inc.
- Ghemical – The Ghemical computational chemistry package.
- MM1GP – Ghemical molecular mechanics file.
- HIN – IN / HIV files from HyperChem – Hypercube, Inc.
- Jaguar – Schrodinger, LLC.
- MOLPRO – Molpro output.
- MOPAC – MOPAC 93/97/2002 output (public domain).
- MGF – MOPAC 2007 (v.7.101) graphf output (public domain).
- NWCHEM – NWChem output – Pacific Northwest National Laboratory.
- odydata – Odyssey data – WaveFunction, Inc.
- xodydata – Odyssey XML data – WaveFunction, Inc.
- QOUT – Q-Chem, Inc.
- SHELX – Structural Chemistry Department, University of Göttingen (Germany).
- SMOL – Spartan data – Wavefunction, Inc.
- spinput – Spartan data – Wavefunction, Inc.
- GRO – Gromos87 format from GROMACS.
- PQR – Modified pdb format including charge and radius.
- Amber – The Amber package of molecular simulation programs.
- JME – Java Molecular Editor – Peter Ertl.
- CASTEP – The CASTEP software package, uses density functional theory.
- FHI-aims – Full-potential / all-electron electronic structure theory with local orbitals – Fritz-Haber-Institut der Max-Planck-Gesellschaft.
- VASP – VASP / VAMP / Vienna ab-initio simulation package.
- DGrid – Miroslav Kohout, Max-Planck Institute.
- ADF – ADF output – Amsterdam Density Functional.
- XSD – Accelrys Materials Studio.
- AGL – ArgusLab.
- DFT – Wien2k.
- AMPAC – AMPAC output – Semichem, Inc.
- WebMO – WebMO interface to computational chemistry packages.
- Molden – Electron density / molecular orbitals.
- PSI3 – Output files from the PSI3 suite of quantum chemical programs.
- CRYSTAL – Output files from CRYSTAL, a computational tool for solid state chemistry and physics. Theoretical Chemistry Group, Univ. Torino, Italy.
- Animations.
- Vibrations.
- Surfaces.
- Orbitals.
- Support for unit cell and symmetry operations.
- Schematic shapes for secondary structures in biomolecules.
- Measurements:
- Distance.
- Angle.
- Torsion angle.
- Support for the RasMol/Chime scripting language.
- JavaScript support library (Jmol.js).
- Exports to jpg, png, gif, ppm, pdf, POV-Ray, Gaussian, Maya, vrml, x3d, idtf, web page.
- Fully internationalised – Multi-language:
- Translated into multiple languages: Catalan (ca), Chinese (both zh_CN and zh_TW) Czech (cs), Dutch (nl), French (fr), German (de), Hungarian (hu), Italian (it), Korean (ko), Portuguese – Brazil (pt_BR), Spanish (es), Turkish (tr), (in addition to the native American English, en-US, and British English, en-GB).
-
- Automatically adopts the language of the user’s operating system, if it is among the translations available.
Website: jmol.sourceforge.net
Support: Handbook
Developer: Jmol Development Team
License: GNU Lesser General Public License
Jmol is written in Java. Learn Java with our recommended free books and free tutorials.
Related Software
| Chemistry Tools | |
|---|---|
| GROMACS | Versatile package to perform molecular dynamics |
| tomviz | Process, visualize, and analyze 3D tomographic data |
| Psi4 | Ab initio quantum chemistry software |
| NWChem | Ab initio computational chemistry software package |
| PyMOL | OpenGL molecular graphics system written in Python |
| LAMMPS | Classical molecular dynamics simulator |
| CP2K | Atomistic simulations of solid state, liquid, molecular and biological systems |
| RDKit | Cheminformatics and machine-learning software |
| GAMESS | General ab initio quantum chemistry package |
| Avogadro | Advanced molecular editor |
| OpenMM | High-performance toolkit for molecular simulation |
| Gabedit | Graphical user interface to computational chemistry packages |
| Open Babel | Converts and manipulates chemical data files |
| Jmol | Viewer for three-dimensional chemical structures |
| pyscf | Quantum chemistry framework |
| Kalzium | Full-featured chemistry application for KDE 5 |
| Ketcher | Web-based chemical structure editor |
| XDrawChem | 2D editor for chemical structures and reactions |
| Cantera | Chemical kinetics, thermodynamics, and transport tool suite |
| Orac | OpenMP/MPI molecular dynamics engine to simulate solvated biomolecules |
| JChemPaint | Chemical 2D structure editor |
| MPQC | Computes the properties of molecules, ab initio |
| Indigo | Universal cheminformatics toolkit for working with molecules and reactions |
| GPAW | Python package for density-functional theory calculations |
| DFTB+ | General package for performing fast atomistic calculations |
| BKChem | 2D molecule editor written in Python |
| MDynaMix | General purpose molecular dynamics |
| ChemCanvas | 2D chemical drawing tool |
| MoleQueue | Abstract, manage, and coordinate the execution of tasks |
Read our verdict in the software roundup.
 Explore our comprehensive directory of recommended free and open source software. Our carefully curated collection spans every major software category. Explore our comprehensive directory of recommended free and open source software. Our carefully curated collection spans every major software category.This directory is part of our ongoing series of informative articles for Linux enthusiasts. It features hundreds of detailed reviews, along with open source alternatives to proprietary solutions from major corporations such as Google, Microsoft, Apple, Adobe, IBM, Cisco, Oracle, and Autodesk. You’ll also find interesting projects to try, hardware coverage, free programming books and tutorials, and much more. Discovered a useful open source Linux program that we haven’t covered yet? Let us know by completing this form. |

