The UCSC Genome Browser is an on-line, and downloadable, genome browser.
This is an interactive website offering access to genome sequence data from a variety of vertebrate and invertebrate species and major model organisms, integrated with a large collection of aligned annotations.
The Browser is a graphical viewer optimized to support fast interactive performance and is an open-source, web-based tool suite built on top of a MySQL database for rapid visualization, examination, and querying of the data at many levels.
Key Features
- Wide collection of annotation datasets (known as “tracks” and presented graphically), including mRNA alignments, mappings of DNA repeat elements, gene predictions, gene-expression data, disease-association data (representing the relationships of genes to diseases), and mappings of commercially available gene chips (e.g. Illumina and Agilent).
- Instant access to the alignments of any RNA to any of the hosted species
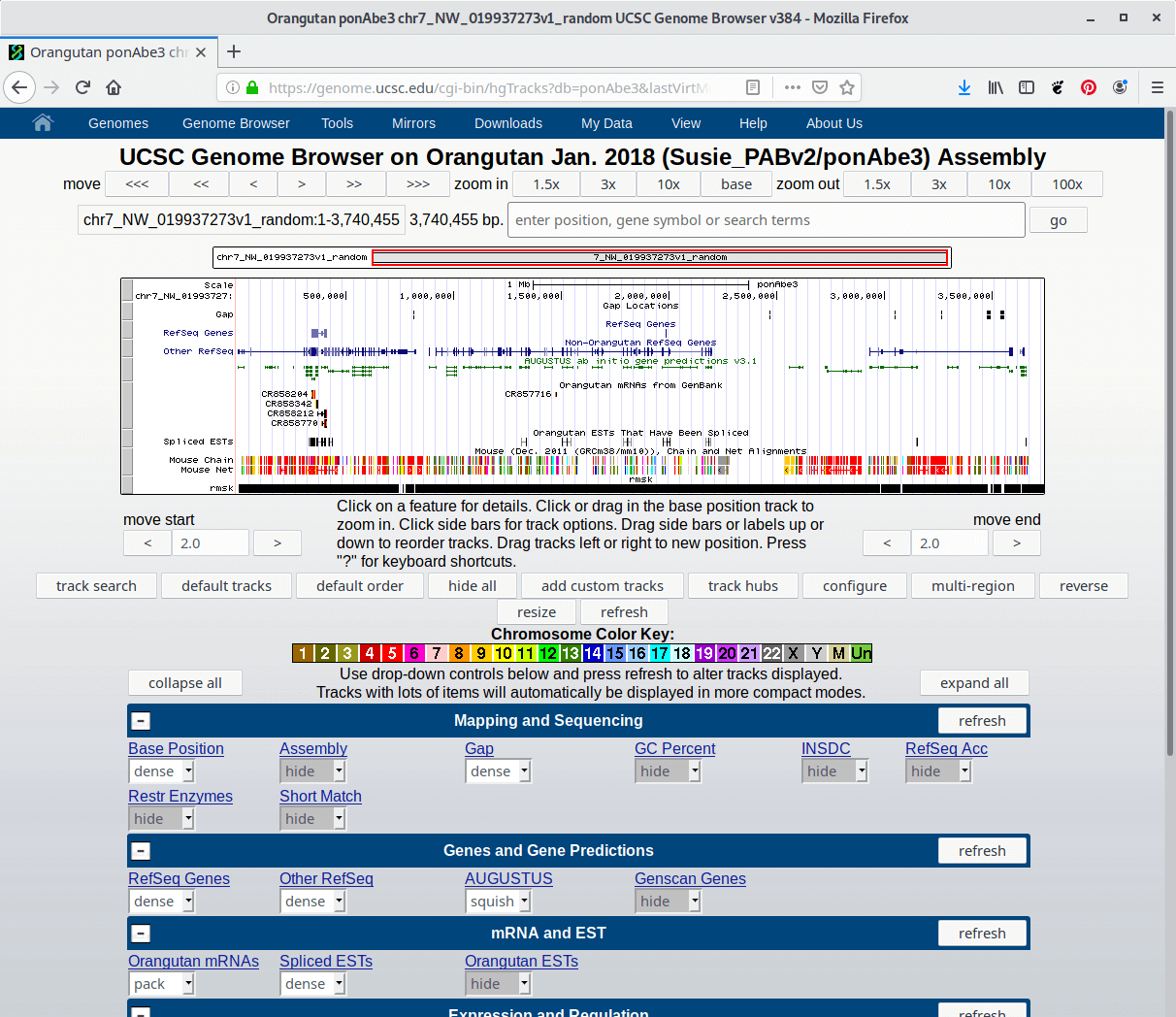
- Continuously variable nature of the display. Sequence of any size can be displayed, from a single DNA base up to the entire chromosome (human chr1 = 245 million bases, Mb) with full annotation tracks.
- Highly configurable.
Website: genome.ucsc.edu
Support: Blog
Developer: UC Santa Cruz
License: UCSC Browser code base is open-source for non-commercial use. A license is required for commercial use.
Related Software
| Web-Based Desktop Genome Browsers | |
|---|---|
| Ensembl | Resource for geneticists, molecular biologists and other researchers |
| Genome Browser | Interactively visualize genomic data |
| GDV | Exploration and analysis of eukaryotic RefSeq genome assemblies |
| HiGlass | Explore and compare genomic contact matrices and tracks |
| igv.js | Embeddable genomic visualization |
| NGB | Web-based NGS data viewer |
| JBrowse 2 | Modern React-based genome browser |
| trackplot | Visualize various next-generation sequencing data |
| GIVE | Genomic Interactive Visualization Engine |
| Genoverse | HTML5 scrollable genome browser |
| Epigenome Browser | Visualization, integration and analysis tools for epigenomic datasets |
Read our verdict in the software roundup.
 Explore our comprehensive directory of recommended free and open source software. Our carefully curated collection spans every major software category. Explore our comprehensive directory of recommended free and open source software. Our carefully curated collection spans every major software category.This directory is part of our ongoing series of informative articles for Linux enthusiasts. It features hundreds of detailed reviews, along with open source alternatives to proprietary solutions from major corporations such as Google, Microsoft, Apple, Adobe, IBM, Cisco, Oracle, and Autodesk. You’ll also find interesting projects to try, hardware coverage, free programming books and tutorials, and much more. Discovered a useful open source Linux program that we haven’t covered yet? Let us know by completing this form. |

