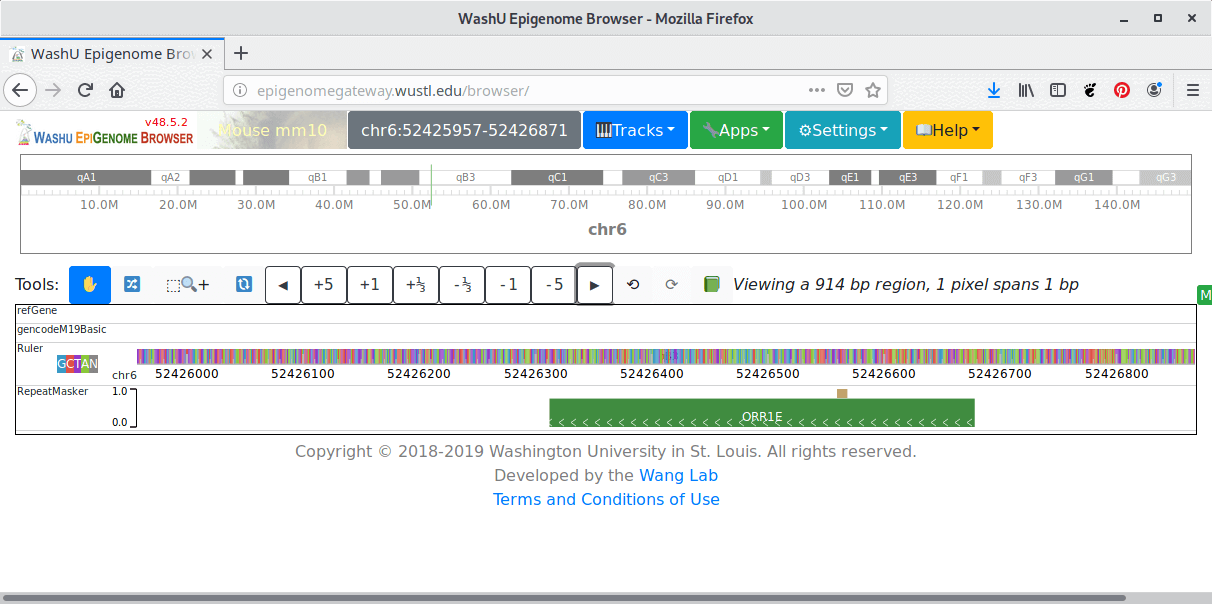
The WashU Epigenome Browser provides visualization, integration and analysis tools for epigenomic datasets.
It has provided the scientific community with data from large consortia including the Roadmap Epigenomics and the ENCODE projects.
The browser is also available as Docker images,
Features include:
- Responsive user interface.
- Undo/Redo functionality.
- Visualization using virtual reality (VR), which has implications in biology education and the study of 3D chromatin structure; (ii)
- Expanded public data hubs, including data from the 4DN, ENCODE, Roadmap Epigenomics, TaRGET, IHEC and TCGA consortia.
- History of interactions, which enables undo and redo
- Live Browsing, which allows multiple users to collaborate remotely on the same session. Share screen with your PI, collaborators and friends.
- Ability to visualize local tracks and data hubs.
Website: epigenomegateway.wustl.edu
Support: Documentation, Video Tutorials, GitHub Code Repository
Developer: Wang Lab – Department of Genetics and Center for Genome Sciences & Systems Biology at Washington University in St. Louis, School of Medicine
License: Open Source
WashU Epigenome Browser is written in JavaScript and C. Learn JavaScript with our recommended free books and free tutorials. Learn C with our recommended free books and free tutorials.
Return to Web Based Genome Browsers
| Popular series | |
|---|---|
| The largest compilation of the best free and open source software in the universe. Each article is supplied with a legendary ratings chart helping you to make informed decisions. | |
| Hundreds of in-depth reviews offering our unbiased and expert opinion on software. We offer helpful and impartial information. | |
| The Big List of Active Linux Distros is a large compilation of actively developed Linux distributions. | |
| Replace proprietary software with open source alternatives: Google, Microsoft, Apple, Adobe, IBM, Autodesk, Oracle, Atlassian, Corel, Cisco, Intuit, SAS, Progress, Salesforce, and Citrix | |
| Awesome Free Linux Games Tools showcases a series of tools that making gaming on Linux a more pleasurable experience. This is a new series. | |
| Machine Learning explores practical applications of machine learning and deep learning from a Linux perspective. We've written reviews of more than 40 self-hosted apps. All are free and open source. | |
| New to Linux? Read our Linux for Starters series. We start right at the basics and teach you everything you need to know to get started with Linux. | |
| Alternatives to popular CLI tools showcases essential tools that are modern replacements for core Linux utilities. | |
| Essential Linux system tools focuses on small, indispensable utilities, useful for system administrators as well as regular users. | |
| Linux utilities to maximise your productivity. Small, indispensable tools, useful for anyone running a Linux machine. | |
| Surveys popular streaming services from a Linux perspective: Amazon Music Unlimited, Myuzi, Spotify, Deezer, Tidal. | |
| Saving Money with Linux looks at how you can reduce your energy bills running Linux. | |
| Home computers became commonplace in the 1980s. Emulate home computers including the Commodore 64, Amiga, Atari ST, ZX81, Amstrad CPC, and ZX Spectrum. | |
| Now and Then examines how promising open source software fared over the years. It can be a bumpy ride. | |
| Linux at Home looks at a range of home activities where Linux can play its part, making the most of our time at home, keeping active and engaged. | |
| Linux Candy reveals the lighter side of Linux. Have some fun and escape from the daily drudgery. | |
| Getting Started with Docker helps you master Docker, a set of platform as a service products that delivers software in packages called containers. | |
| Best Free Android Apps. We showcase free Android apps that are definitely worth downloading. There's a strict eligibility criteria for inclusion in this series. | |
| These best free books accelerate your learning of every programming language. Learn a new language today! | |
| These free tutorials offer the perfect tonic to our free programming books series. | |
| Linux Around The World showcases usergroups that are relevant to Linux enthusiasts. Great ways to meet up with fellow enthusiasts. | |
| Stars and Stripes is an occasional series looking at the impact of Linux in the USA. | |

