GNomEx (Genomic LIMS and Data Repository) is an open-source genomics LIMS and analysis project center for organizing, annotating, tracking, and distributing raw genomic data and associated downstream analysis.
This application holds annotated experiments and downstream analysis and serves data tracks to popular genome browsers such as IGB, IGV, and UCSC genome browser. The LIMS handles all aspects of the experiment from order through results delivery.
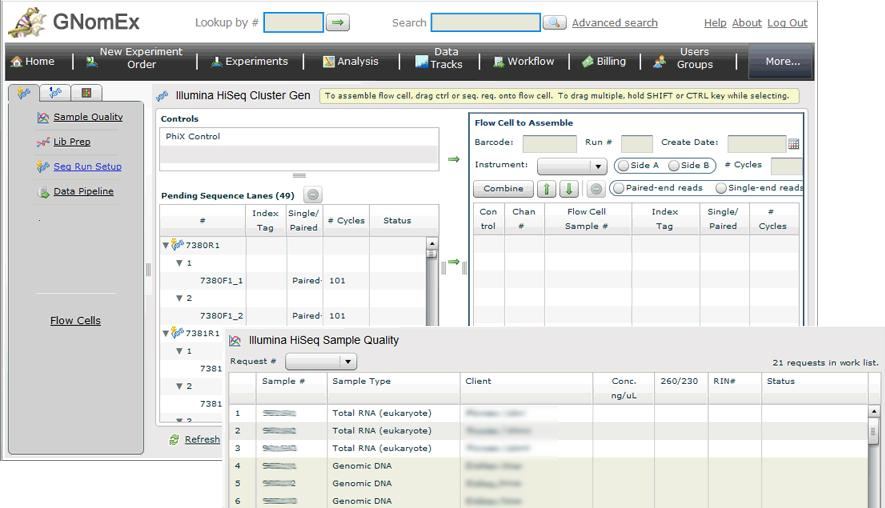
Experiment platforms supported include Illumina HiSeq, MiSeq, iScan, ABI Sanger sequencing, Affy and Agilent Microarrays, Sequenom MassArray and Bioanalyzer.
Key Features
- Submit an Experiment Order to a Core Facility and receive emails notifying you of progress.
- Create an experiment and upload your own raw data files.
- Fully annotate your experiment for each submitted sample.
- Register downstream analysis performed on the Experiment, tracking all of the analysis pipeline details relevant to the data, for example, the Genome Build the sequence was aligned to.
- Download the experiment results (raw data files) as well as analyzed data sets.
- Organize analyzed data sets that can be viewed in a Genome Browser.
- Organize experiments and analyses under Topics to aggregate data into unified view.
- Dial in the access to your data – owner, lab members, institution, or public. Also, grant access to specific collaborators.
- Search the repository for a key term or perform more advanced criteria based searches.
- Configure different Experiment platforms based on the major categories of Next Gen Sequencing (Illumina HiSeq and MiSeq), Microarray, iScan, Capillary Sequencing, Sample Quality, Fragment Analysis, Mitochondrial DLoop Sequencing, and Plate Cherry Picking.
- Configure annotations required for experiment samples and analyses. Handle wide range of annotations using text, checkbox, dropdowns, multi-select lists, and URLs.
- For core facilities, manage experiment orders through the workflow and automate your billing.
- For administrators, manage user accounts and labs.
- Web tools to browse, search, modify, and download data files associated with datasets and meta-data.
Website: github.com/hci-gnomex/gnomex
Support:
Developer: David A. Nix, Tonya L. Di Sera, Brett A. Milash, and Samir J. Courdy
License: GNU General Public License v3.0
GNomEx is written in Java. Learn Java with our recommended free books and free tutorials.
Related Software
| Laboratory Information Management System | |
|---|---|
| eLabFTW | Electronic lab notebook manager for research teams |
| SENAITE | Enterprise Open Source Laboratory System |
| OpenMRS | Enterprise electronic medical record system platform |
| Bika LIMS | Powerful, flexible web based LIMS with full content management integration |
| Occhiolino | Modern Laboratory Information Management System |
| C4G BLIS | Track patients, specimens and laboratory results |
| OpenELIS | Designed for resource-constrained laboratories |
| FreeLIMS | Lets users perform Bio-Informatics |
| MetaLIMS | LIMS for small metagenomic laboratories |
| GNomEx | Genomic LIMS and Data Repository |
| Baobab LIMS | LIMS for biobanking |
| Open-LIMS | Management for projects, samples and their related data |
Read our verdict in the software roundup.
 Explore our comprehensive directory of recommended free and open source software. Our carefully curated collection spans every major software category. Explore our comprehensive directory of recommended free and open source software. Our carefully curated collection spans every major software category.This directory is part of our ongoing series of informative articles for Linux enthusiasts. It features hundreds of detailed reviews, along with open source alternatives to proprietary solutions from major corporations such as Google, Microsoft, Apple, Adobe, IBM, Cisco, Oracle, and Autodesk. You’ll also find interesting projects to try, hardware coverage, free programming books and tutorials, and much more. Discovered a useful open source Linux program that we haven’t covered yet? Let us know by completing this form. |

