GROMACS is a molecular dynamics simulator, with building and analysis tools.
GROMACS is a versatile package to perform molecular dynamics, i.e. simulate the Newtonian equations of motion for systems with hundreds to millions of particles.
It is primarily designed for biochemical molecules like proteins and lipids that have a lot of complicated bonded interactions, but since GROMACS is extremely fast at calculating the nonbonded interactions (that usually dominate simulations) many groups are also using it for research on non-biological systems, e.g. polymers.
It provides extremely high performance compared to other competitors. A lot of algorithmic optimizations have been introduced in the code. The software is normally 3-10 times faster than any other program.
There is ongoing development to extend GROMACS with interfaces both to Quantum Chemistry and Bioinformatics/databases.
Key Features
- User-friendly, with topologies and parameter files written in clear text format. There is a lot of consistency checking, and clear error messages are issued when something is wrong.
- There is no scripting language – all programs use a simple interface with command line options for input and output files.
- Extensive manuals provided free of charge in electronic or paper format.
- Integrated graphical user interface available for all programs
- As the simulation is proceeding, GROMACS will continuously tell you how far it has come, and what time and date it expects to be finished.
- Both run input files and trajectories are independent of hardware endian and can thus be read by any version GROMACS, even if it was compiled using a different floating-point precision. All files from GROMACS 2.0 can further be used in the new version 3.
- Write coordinates using lossy compression, which provides a very compact way of storing trajectory data. The accuracy can be selected by the user.
- Large selection of flexible tools for trajectory analysis – you won’t have to write any code to perform routine analyses. The output is further provided in the form of finished Xmgr/Grace graphs, with axis labels, legends, etc. already in place.
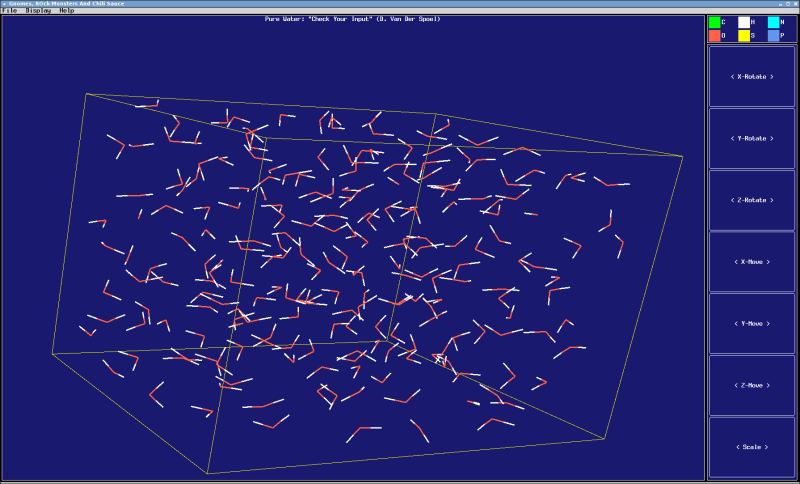
- A basic trajectory viewer that only requires standard X libraries is included, and several external visualization tools can read the GROMACS file formats.
- Can be run in parallel, using standard MPI communication.
- Contains several state-of-the-art algorithms that make it possible to extend the time steps is simulations significantly, and thereby further enhance performance without sacrificing accuracy or detail.
- Includes a fully automated topology builder for proteins, even multimeric structures. Building blocks are available for the 20 standard aminoacid residues as well as some modified ones, the 4 nucleotide and 4 deoxinucleotide resides, several sugars and lipids, and some special groups like hemes and several small molecules.
Website: www.gromacs.org
Support: Tutorial
Developer: David van der Spoel, Berk Hess, Erik Lindahl, and many contributors
License: GNU General Public License v2.0
GROMACS is written in C++ and C. Learn C++ with our recommended free books and free tutorials. Learn C with our recommended free books and free tutorials.
Related Software
| Biology Tools | |
|---|---|
| EMBOSS | The European Molecular Biology Open Software Suite |
| NAMD | Parallel, object-oriented molecular dynamics |
| GROMACS | Molecular dynamics simulator, with building and analysis tools |
| VMD | Displays, animates, and analyzes large biomolecular systems 3-D graphics |
| simuPOP | Forward-time population genetics simulation environment |
| MUSCLE | MUltiple Sequence Comparison by Log-Expectation |
| SeaView | Graphical user interface for molecular phyologeny |
| SeqViz | DNA, RNA, and protein sequence viewer |
| TREE-PUZZLE | Reconstruction of phylogenetic trees by maximum likelihood |
| TreeView X | Displays and prints phylogenetic trees |
Read our verdict in the software roundup.
| Chemistry Tools | |
|---|---|
| GROMACS | Versatile package to perform molecular dynamics |
| tomviz | Process, visualize, and analyze 3D tomographic data |
| Psi4 | Ab initio quantum chemistry software |
| NWChem | Ab initio computational chemistry software package |
| PyMOL | OpenGL molecular graphics system written in Python |
| LAMMPS | Classical molecular dynamics simulator |
| CP2K | Atomistic simulations of solid state, liquid, molecular and biological systems |
| RDKit | Cheminformatics and machine-learning software |
| GAMESS | General ab initio quantum chemistry package |
| Avogadro | Advanced molecular editor |
| OpenMM | High-performance toolkit for molecular simulation |
| Gabedit | Graphical user interface to computational chemistry packages |
| Open Babel | Converts and manipulates chemical data files |
| Jmol | Viewer for three-dimensional chemical structures |
| pyscf | Quantum chemistry framework |
| Kalzium | Full-featured chemistry application for KDE 5 |
| Ketcher | Web-based chemical structure editor |
| XDrawChem | 2D editor for chemical structures and reactions |
| Cantera | Chemical kinetics, thermodynamics, and transport tool suite |
| Orac | OpenMP/MPI molecular dynamics engine to simulate solvated biomolecules |
| JChemPaint | Chemical 2D structure editor |
| MPQC | Computes the properties of molecules, ab initio |
| Indigo | Universal cheminformatics toolkit for working with molecules and reactions |
| GPAW | Python package for density-functional theory calculations |
| DFTB+ | General package for performing fast atomistic calculations |
| BKChem | 2D molecule editor written in Python |
| MDynaMix | General purpose molecular dynamics |
| ChemCanvas | 2D chemical drawing tool |
| MoleQueue | Abstract, manage, and coordinate the execution of tasks |
Read our verdict in the software roundup.
| Bioinformatics Tools | |
|---|---|
| Bioconductor | Analysis and comprehension of high-throughput genomic data |
| Biopython | Tools for biological computation written in Python |
| UGENE | Set of integrated bioinformatics software |
| BioPerl | Perl tools for computational molecular biology |
| GROMACS | Versatile package to perform molecular dynamics |
| IGV | High-performance visualization genome browser tool |
| GATK | Genomic analysis toolkit focused on variant discovery |
| BioJava | Provides Java tools for processing biological data |
| InterMine | Integrate biological data sources |
| bedtools | Powerful toolset for genome arithmetic |
| EMBOSS | The European Molecular Biology Open Software Suite |
| BLAST | Algorithm for comparing primary biological sequence information |
| Galaxy | Web-based platform for data-intensive computational research |
| minimap2 | Versatile sequence alignment program |
| Jalview | Multiple sequence alignment editing, visualisation and analysis |
| samtools | Manipulate next-generation sequencing data |
| BCFtools | Variant calling and manipulating files in the Variant Call Format |
| FastQC | Quality control tool for high throughput sequence data |
| SPAdes | Versatile toolkit for assembling and analysing sequencing data |
| GenomeTools | Collection of bioinformatics tools |
| AliView | Alignment viewer and editor |
| mothur | Analyze microbial communities |
| Bandage | Visualising de novo assembly graphs |
| cramino | BAM/CRAM quality evaluation |
| abPOA | Adaptive banded Partial Order Alignment |
| Taverna Workbench | For designing and executing bioinformatics workflows |
| geWorkbench | Software platform for integrated genomic data analysis |
| Bioclipse | Rich-client platform chemistry and biology workbench |
Read our verdict in the software roundup.
 Explore our comprehensive directory of recommended free and open source software. Our carefully curated collection spans every major software category. Explore our comprehensive directory of recommended free and open source software. Our carefully curated collection spans every major software category.This directory is part of our ongoing series of informative articles for Linux enthusiasts. It features hundreds of detailed reviews, along with open source alternatives to proprietary solutions from major corporations such as Google, Microsoft, Apple, Adobe, IBM, Cisco, Oracle, and Autodesk. You’ll also find interesting projects to try, hardware coverage, free programming books and tutorials, and much more. Discovered a useful open source Linux program that we haven’t covered yet? Let us know by completing this form. |

